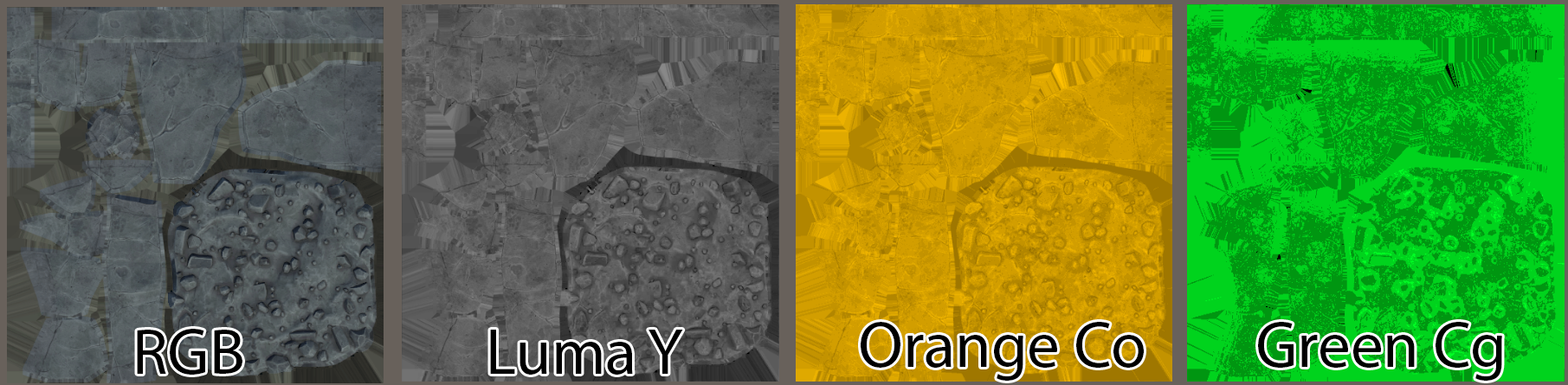
XYZ Spectral Sensitivity Curves
First, let’s get those Gaussian curves for the XYZ spectral sensitivity curves. I’ve pre-generated these into (you guessed it, CSV).
Put this into your development directory, with everything else:
In our __init__ function, do the following:
cmf = open("cie-cmf.csv", "r")
contents = cmf.readlines()
CIE_CMF = [None] * 42
for line in contents:
data = line .rstrip("\n").split(",")
CIE_CMF[int((int(data [0]) - 380) / 10)] = [float(data [1]),float(data [2]),float(data [3])]
cmf.close()
for nm in range(380,790,10):
self.CIE_XYZ_Spectral_Sensitivity_Curve[nm] = CIE_CMF[int((nm - 380)/10)]And declare at the top of our class, CIE_CMF:
class SkinAbsorption:
O2Hb = {}
Hb = {}
CIE_XYZ_Spectral_Sensitivity_Curve = {}
CIE_XYZ_Spectral_Sensitivity_Curve = {}
This will store the values in a dictionary.
RGB Matrices
Now to add in our RGB matrix and gamma correction for sRGB:
def gamma_correction(self, C):
abs_C = abs(C)
if abs_C > 0.0031308:
return 1.055 * pow(abs_C,1/2.4) - 0.055
else:
return 12.92 * C
def XYZ_to_sRGB(self, xyz):
x = xyz[0]/10
y = xyz[1]/10
z = xyz[2]/10
mat3x3 = [(3.2406, -1.5372, -0.4986), (-0.9689, 1.8758, 0.0415), (0.0557, -0.204, 1.057)]
r = self.gamma_correction(x * mat3x3[0][0] + y * mat3x3[0][1] + z * mat3x3[0][2])
g = self.gamma_correction(x * mat3x3[1][0] + y * mat3x3[1][1] + z * mat3x3[1][2])
b = self.gamma_correction(x * mat3x3[2][0] + y * mat3x3[2][1] + z * mat3x3[2][2])
sRGB = (int(r*255),int(g*255),int(b*255)) #needs to be 0 - 255 for outputing to color image
return sRGBConverting Our Data
Our data isn’t in a usable state yet. We need to restructure the data coming from the Monte Carlo simulation to be organised by melanin blend, melanin fraction, hemoglobin fraction – this is so we can output a pixel of colour for each value.
We’ll replace all our CSV output code, and instead of returning a string, we’ll return tuples of the values, like so:
def GetReflectanceValues(self, Cm, Ch, Bm):
wavelengths = range(380,790,10)
reflectances = []
for nm in wavelengths:
# First layer absorption - Epidermis
SAV_eumelanin_L = 6.6 * pow(10,10) * pow(nm,-3.33)
SAV_pheomelanin_L = 2.9 * pow(10,14) * pow(nm,-4.75)
epidermal_hemoglobin_fraction = Ch * 0.25
# baseline - used in both layers
baselineSkinAbsorption_L = 0.0244 + 8.54 * pow(10,-(nm-220)/123)
# Epidermis Absorption Coefficient:
epidermis = Cm * ((Bm * SAV_eumelanin_L) +((1 - Bm ) * SAV_pheomelanin_L)) + ((1 - epidermal_hemoglobin_fraction) * baselineSkinAbsorption_L)
# Second layer absorption - Dermis
gammaOxy = self.O2Hb[int(nm)]
gammaDeoxy = self.Hb[int(nm)]
A = 0.75 * gammaOxy + (1 - 0.75) * gammaDeoxy
dermis = Ch * (A + (( 1 - Ch) * baselineSkinAbsorption_L))
# Scattering coefficients
scattering_epidermis = pow(14.74 * nm, -0.22) + 2.2 * pow(10,11) * pow(nm, -4)
scattering_dermis = scattering_epidermis * 0.5
reflectance = self.MonteCarlo(epidermis, scattering_epidermis, dermis, scattering_dermis, nm)
reflectances.append( (reflectance,Cm,Ch,Bm,nm,epidermis,dermis,baselineSkinAbsorption_L))
return reflectances
And adjust the generate function:
def Generate(self):
groupByBlend = []
for Bm in [0.01,0.5,0.99]:
groupByMelanin = []
for Cm in [0.002,0.0135,0.0425,0.1,0.185,0.32,0.5]:
groupByHemoglobin = []
for Ch in [0.003,0.02,0.07,0.16,0.32]:
values = self.GetReflectanceValues( Cm, Ch, Bm )
groupByHemoglobin.append(values)
groupByMelanin.append(groupByHemoglobin)
groupByBlend.append(groupByMelanin)
return groupByBlendNow, above the return statement add the following to convert all our data to sRGB:
XYZ_Colors = []
specular_col = []
for melanin_blend in groupByBlend:
for melanin_fraction in melanin_blend :
for hemoglobin_fraction in melanin_fraction:
total = (0,0,0)
spec = [0.0] * 41
for data in hemoglobin_fraction:
reflectance = data[0]
spec[int((data[4] -380) / 10)] = data[0]
xyz = self.CIE_XYZ_Spectral_Sensitivity_Curve.get(data[4])
x = xyz[0] * reflectance
y = xyz[1] * reflectance
z = xyz[2] * reflectance
total = (total[0] + x, total[1] + y, total[2] + z)
XYZ_Colors.append(total)
pixelsRGB = []
for xyz in XYZ_Colors:
pixelsRGB.append(self.XYZ_to_sRGB(xyz))Next step, we’ll make an image!
For reference, current code:
import random
import math
from PIL import Image
class SkinAbsorption:
O2Hb = {}
Hb = {}
CIE_XYZ_Spectral_Sensitivity_Curve = {}
def __init__(self):
print("init SkinAbsorption")
cmf = open("cie-cmf.csv", "r")
contents = cmf.readlines()
CIE_CMF = [None] * 42
for line in contents:
data = line .rstrip("\n").split(",")
CIE_CMF[int((int(data [0]) - 380) / 10)] = [float(data [1]),float(data [2]),float(data [3])]
cmf.close()
for nm in range(380,790,10):
self.CIE_XYZ_Spectral_Sensitivity_Curve[nm] = CIE_CMF[int((nm - 380)/10)]
HbFile = open('hb.csv', "r")
O2HbFile = open('O2Hb.csv', "r")
HbLines = HbFile.readlines()
for line in HbLines:
splitLine = line.split(",")
self.Hb[int(splitLine[0])] = float(splitLine[1].rstrip("\n"))
O2HbLines = O2HbFile.readlines()
for line in O2HbLines:
splitLine = line.split(",")
self.O2Hb[int(splitLine[0])] = float(splitLine[1].rstrip("\n"))
HbFile.close()
O2HbFile.close()
def Generate(self):
groupByBlend = []
for Bm in [0.01,0.5,0.99]:
groupByMelanin = []
for Cm in [0.002,0.0135,0.0425,0.1,0.185,0.32,0.5]:
groupByHemoglobin = []
for Ch in [0.003,0.02,0.07,0.16,0.32]:
values = self.GetReflectanceValues( Cm, Ch, Bm )
groupByHemoglobin.append(values)
groupByMelanin.append(groupByHemoglobin)
groupByBlend.append(groupByMelanin)
XYZ_Colors = []
specular_col = []
for melanin_blend in groupByBlend: # Mb
for melanin_fraction in melanin_blend:
for hemoglobin_fraction in melanin_fraction: # Mf
total = (0,0,0)
spec = [0.0] * 41
for data in hemoglobin_fraction: # Hf
print(data)
reflectance = data[0]
spec[int((data[4] -380) / 10)] = data[0]
xyz = self.CIE_XYZ_Spectral_Sensitivity_Curve.get(data[4])
x = xyz[0] * reflectance
y = xyz[1] * reflectance
z = xyz[2] * reflectance
total = (total[0] + x, total[1] + y, total[2] + z)
XYZ_Colors.append(total)
pixelsRGB = []
for xyz in XYZ_Colors:
pixelsRGB.append(self.XYZ_to_sRGB(xyz))
return pixelsRGB
def GetReflectanceValues(self, Cm, Ch, Bm):
wavelengths = range(380,790,10)
reflectances = []
for nm in wavelengths:
# First layer absorption - Epidermis
SAV_eumelanin_L = 6.6 * pow(10,10) * pow(nm,-3.33)
SAV_pheomelanin_L = 2.9 * pow(10,14) * pow(nm,-4.75)
epidermal_hemoglobin_fraction = Ch * 0.25
# baseline - used in both layers
baselineSkinAbsorption_L = 0.0244 + 8.54 * pow(10,-(nm-220)/123)
# Epidermis Absorption Coefficient:
epidermis = Cm * ((Bm * SAV_eumelanin_L) +((1 - Bm ) * SAV_pheomelanin_L)) + ((1 - epidermal_hemoglobin_fraction) * baselineSkinAbsorption_L)
# Second layer absorption - Dermis
gammaOxy = self.O2Hb[int(nm)]
gammaDeoxy = self.Hb[int(nm)]
A = 0.75 * gammaOxy + (1 - 0.75) * gammaDeoxy
dermis = Ch * (A + (( 1 - Ch) * baselineSkinAbsorption_L))
# Scattering coefficients
scattering_epidermis = pow(14.74 * nm, -0.22) + 2.2 * pow(10,11) * pow(nm, -4)
scattering_dermis = scattering_epidermis * 0.5
reflectance = self.MonteCarlo(epidermis, scattering_epidermis, dermis, scattering_dermis, nm)
reflectances.append((reflectance,Cm,Ch,Bm,nm,epidermis,dermis,baselineSkinAbsorption_L))
return reflectances
def MonteCarlo (self, epi_mua, epi_mus, derm_mua, derm_mus, nm):
# These are our Monte Carlo Light Transport Variables that don't change
Nbins = 1000
Nbinsp1 = 1001
PI = 3.1415926
LIGHTSPEED = 2.997925 * pow(10,10)
ALIVE = 1
DEAD = 0
THRESHOLD = 0.01
CHANCE = 0.1
COS90D = 1 * pow(10,-6)
ONE_MINUS_COSZERO = 1 * pow(10,-12)
COSZERO = 1.0 - 1.0e-12 # cosine of about 1e-6 rad
g = 0.9
nt = 1.33 # Index of refraction
epidermis_thickness = 0.25
x = 0.0
y = 0.0
z = 0.0 # photon position
ux = 0.0
uy = 0.0
uz = 0.0 # photon trajectory as cosines
uxx = 0.0
uyy = 0.0
uzz = 0.0 # temporary values used during SPIN
s = 0.0 # step sizes. s = -log(RND)/mus [cm]
costheta = 0.0 # cos(theta)
sintheta = 0.0 # sin(theta)
cospsi = 0.0 # cos(psi)
sinpsi = 0.0 # sin(psi)
psi = 0.0 # azimuthal angle
i_photon = 0.0 # current photon
W = 0.0 # photon weight
absorb = 0.0 # weighted deposited in a step due to absorption
photon_status = 0.0 # flag = ALIVE=1 or DEAD=0
ReflBin = [None] * Nbinsp1 #bin to store weights of escaped photos for reflectivity
epi_albedo = epi_mus/(epi_mus + epi_mua) # albedo of tissue
derm_albedo = derm_mus/(derm_mus + derm_mua) # albedo of tissue
Nphotons = 1000 # number of photons in simulation
NR = Nbins # number of radial positions
radial_size = 2.5 # maximum radial size
r = 0.0 # radial position
dr = radial_size/NR; # cm, radial bin size
ir = 0 # index to radial position
shellvolume = 0.0 # volume of shell at radial position r
CNT = 0.0 # total count of photon weight summed over all bins
rnd = 0.0 # assigned random value 0-1
u = 0.0
temp = 0.0 # dummy variables
# Inits
random.seed(0)
RandomNum = random.random()
for i in range(NR+1):
ReflBin[i] = 0
while True:
i_photon = i_photon + 1
W = 1.0
photon_status = ALIVE
x= 0
y = 0
z = 0
#Randomly set photon trajectory to yield an isotropic source.
costheta = 2.0 * random.random() - 1.0
sintheta = math.sqrt(1.0 - costheta*costheta)
psi = 2.0 * PI * random.random()
ux = sintheta * math.cos(psi)
uy = sintheta * math.sin(psi)
uz = (abs(costheta)) # on the first step we want to head down, into the tissue, so > 0
# Propagate one photon until it dies as determined by ROULETTE.
# or if it reaches the surface again
it = 0
max_iterations = 100000 # to help avoid infinite loops in case we do something wrong
# we'll hit epidermis first, so set mua/mus to those scattering/absorption values
mua = epi_mua
mus = epi_mus
albedo = epi_albedo
while True:
it = it + 1
rnd = random.random()
while rnd <= 0.0: # make sure it is > 0.0
rnd = random.random()
s = -math.log(rnd)/(mua + mus)
x = x + (s * ux)
y = y + (s * uy)
z = z + (s * uz)
if uz < 0:
# calculate partial step to reach boundary surface
s1 = abs(z/uz)
# move back
x = x - (s * ux)
y = y - (s * uy)
z = z - (s * uz)
# take partial step
x = x + (s1 * ux)
y = y + (s1 * uy)
z = z + (s1 * uz)
# photon is now at the surface boundary, figure out how much escaped and how much was reflected
internal_reflectance = self.RFresnel(1.0,nt, -uz )
#Add weighted reflectance of escpaed photon to reflectance bin
external_reflectance = 1 - internal_reflectance
r = math.sqrt(x*x + y*y)
ir = (r/dr)
if ir >= NR:
ir = NR
ReflBin[int(ir)] = ReflBin[int(ir)] + (W * external_reflectance)
# Bounce the photon back into the skin
W = internal_reflectance * W
uz = -uz
x = (s-s1) * ux
y = (s-s1) * uy
z = (s-s1) * uz
# check if we have passed into the second layer, or the first
if z <= epidermis_thickness:
mua = epi_mua
mus = epi_mus
albedo = epi_albedo
else:
mua = derm_mua
mus = derm_mus
albedo = derm_albedo
''' DROP '''
absorb = W*(1 - albedo)
W = W - absorb
''' SPIN '''
# Sample for costheta
rnd = random.random()
if (g == 0.0):
costheta = 2.0*rnd - 1.0
else:
temp = (1.0 - g*g)/(1.0 - g + 2*g*rnd)
costheta = (1.0 + g*g - temp*temp)/(2.0*g)
sintheta = math.sqrt(1.0 - costheta*costheta)
# Sample psi.
psi = 2.0*PI*random.random()
cospsi = math.cos(psi)
if (psi < PI):
sinpsi = math.sqrt(1.0 - cospsi*cospsi)
else:
sinpsi = -math.sqrt(1.0 - cospsi*cospsi)
# New trajectory.
if (1 - abs(uz) <= ONE_MINUS_COSZERO) : # close to perpendicular.
uxx = sintheta * cospsi
uyy = sintheta * sinpsi
uzz = costheta * self.SIGN(uz) # SIGN() is faster than division.
else: # usually use this option
temp = math.sqrt(1.0 - uz * uz)
uxx = sintheta * (ux * uz * cospsi - uy * sinpsi) / temp + ux * costheta
uyy = sintheta * (uy * uz * cospsi + ux * sinpsi) / temp + uy * costheta
uzz = -sintheta * cospsi * temp + uz * costheta
# Update trajectory
ux = uxx
uy = uyy
uz = uzz
# Check Roulette
if (W < THRESHOLD):
if (random.random() <= CHANCE):
W = W / CHANCE
else:
photon_status = DEAD
if photon_status is DEAD:
break
if it > max_iterations:
break
if i_photon >= Nphotons:
break
total_reflection = 0.0
for each in range(NR+1):
total_reflection = total_reflection + ReflBin[each]/Nphotons
return total_reflection
def RFresnel(self, n1, n2, cosT1):
r = 0.0
cosT2 = 0.0
COSZERO = 1.0 - 1.0e-12
COS90D = 1 * pow(10,-6)
if n1 == n2: #matched boundary
r = 0.0
cosT2 = cosT1
elif cosT1 > COSZERO: # normal incident
cosT2 = ca1
r = (n2-n1)/(n2+n1)
r *= r
elif cosT1 < COS90D: # very slant
cosT2 = 0.0
r = 1.0
else: #general
sinT1 = math.sqrt(1 - cosT1*cosT1)
sinT2 = n1 * sinT1/n2
if sinT2 >= 1.0:
r = 1.0
cosT2 = 0.0
else:
cosT2 = math.sqrt(1 - sinT2 * sinT2)
cosAP = cosT1*cosT2 - sinT1*sinT2
cosAM = cosT1*cosT2 + sinT1*sinT2
sinAP = sinT1*cosT2 + cosT1*sinT2
sinAM = sinT1*cosT2 - cosT1*sinT2
r = 0.5 * sinAM * sinAM*(cosAM*cosAM+cosAP*cosAP)/(sinAP*sinAP*cosAM*cosAM)
return r
def SIGN(self, x):
if x >=0:
return 1
else:
return 0
def gamma_correction(self, C):
abs_C = abs(C)
if abs_C > 0.0031308:
return 1.055 * pow(abs_C,1/2.4) - 0.055
else:
return 12.92 * C
def XYZ_to_sRGB(self, xyz):
x = xyz[0]/10
y = xyz[1]/10
z = xyz[2]/10
mat3x3 = [(3.2406, -1.5372, -0.4986), (-0.9689, 1.8758, 0.0415), (0.0557, -0.204, 1.057)]
r = self.gamma_correction(x * mat3x3[0][0] + y * mat3x3[0][1] + z * mat3x3[0][2])
g = self.gamma_correction(x * mat3x3[1][0] + y * mat3x3[1][1] + z * mat3x3[1][2])
b = self.gamma_correction(x * mat3x3[2][0] + y * mat3x3[2][1] + z * mat3x3[2][2])
sRGB = (int(r*255),int(g*255),int(b*255)) #needs to be 0 - 255 for outputing to color image
return sRGB
skinAbsorption = SkinAbsorption()
reflectanceValues = skinAbsorption.Generate()